Laboratory Instrumentation, Equipment, and Software
Our laboratory develops and applies advanced analytical methods to answer public-health research questions. Instrumentation, equipment, and custom software in our laboratory that support this research are listed below.
Virtual Tour of the Instrumentation Laboratory
Instrumentation
-
LC-Q/TOF MS: Liquid chromatography coupled to quadrupole/time-of-flight mass spectrometry is used
to perform non-targeted analysis of relatively polar contaminants and metabolites.
-
GCxGC-TOF MS: Comprehensive two dimensional gas chromatography coupled to time-of-flight mass
spectrometry is used to perform non-targeted analysis of relatively non-polar contaminants and
metabolites.
-
LC/MS/MS (x2): Relatively polar molecules including some metabolites are quantified using liquid
chromatography coupled to triple quadrupole mass spectrometry.
-
GC/MS: Gas chromatography coupled to quadrupole mass spectrometry is used to quantify persistent
organic pollutants and other relatively non-polar molecules.
-
LC-ICP MS: Metals are quantified using liquid chromatography coupled to inductively coupled
plasma mass spectrometry.
-
Mercury Analyzer: Mercury is quantified on a specialized instrument.
Sample Preparation Equipment
-
SPE: Solid phase extraction is used to isolate and concentrate molecules of interest from liquid
matrices.
-
PLE: Pressurized liquid extraction isolates molecules of interest from solid/semi-solid matrices.
-
Microwave: Microwave extraction isolates molecules of interest from solid/semi-solid matrices.
-
GPC: Gel permeation chromatography is used to separate molecules of interest from co-extracted
lipids.
-
Large volume nitrogen evaporator: The TurboVap uses a controlled temperature bath and nitrogen
gas vortex to concentrate target analytes in liquid matrices to a desired volume.
-
Small volume nitrogen evaporator: The Reacti-Therm III is a nitrogen blowdown evaporator used to
concentrate molecules of interest in small volumes of liquid matrices, typically prior to instrumental
analysis.
 GCxGC-TOF MS
GCxGC-TOF MS
 LC/MS/MS (1)
LC/MS/MS (1)
 LC/MS/MS (2)
LC/MS/MS (2)
 GC/MS
GC/MS
 LC-ICP MS
LC-ICP MS
 LC-Q/TOF MS
LC-Q/TOF MS
 SPE
SPE
 PLE
PLE
 Microwave
Microwave
 GPC
GPC
 TurboVap
TurboVap
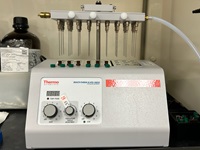 Reacti-Therm
Reacti-Therm
Software
Some of our research projects require the development of custom software to process and summarize data.
Most code is written in R, and is publicly released. For example, custom software is used to:
- Analyze non-targeted mass spectrometry data to assist with compound identification and generate mass
spectral libraries.
- Process quantitative mass spectrometry data to determine target compound concentrations and evaluate
data quality objectives.
Packages
R package for organic/biological mass spectrometry.
Collection of mass spectral libraries.
Instrumentation
- LC-Q/TOF MS: Liquid chromatography coupled to quadrupole/time-of-flight mass spectrometry is used to perform non-targeted analysis of relatively polar contaminants and metabolites.
- GCxGC-TOF MS: Comprehensive two dimensional gas chromatography coupled to time-of-flight mass spectrometry is used to perform non-targeted analysis of relatively non-polar contaminants and metabolites.
- LC/MS/MS (x2): Relatively polar molecules including some metabolites are quantified using liquid chromatography coupled to triple quadrupole mass spectrometry.
- GC/MS: Gas chromatography coupled to quadrupole mass spectrometry is used to quantify persistent organic pollutants and other relatively non-polar molecules.
- LC-ICP MS: Metals are quantified using liquid chromatography coupled to inductively coupled plasma mass spectrometry.
- Mercury Analyzer: Mercury is quantified on a specialized instrument.
Sample Preparation Equipment
- SPE: Solid phase extraction is used to isolate and concentrate molecules of interest from liquid matrices.
- PLE: Pressurized liquid extraction isolates molecules of interest from solid/semi-solid matrices.
- Microwave: Microwave extraction isolates molecules of interest from solid/semi-solid matrices.
- GPC: Gel permeation chromatography is used to separate molecules of interest from co-extracted lipids.
- Large volume nitrogen evaporator: The TurboVap uses a controlled temperature bath and nitrogen gas vortex to concentrate target analytes in liquid matrices to a desired volume.
- Small volume nitrogen evaporator: The Reacti-Therm III is a nitrogen blowdown evaporator used to concentrate molecules of interest in small volumes of liquid matrices, typically prior to instrumental analysis.

GCxGC-TOF MS
LC/MS/MS (1)

LC/MS/MS (2)
GC/MS
LC-ICP MS

LC-Q/TOF MS
SPE

PLE

Microwave
GPC

TurboVap

Reacti-Therm
Software
Some of our research projects require the development of custom software to process and summarize data. Most code is written in R, and is publicly released. For example, custom software is used to:
- Analyze non-targeted mass spectrometry data to assist with compound identification and generate mass spectral libraries.
- Process quantitative mass spectrometry data to determine target compound concentrations and evaluate data quality objectives.
Packages
R package for organic/biological mass spectrometry.
Collection of mass spectral libraries.